Originally trained as an experimental physicist in undergrad, I slowly gravitated to biology – and eventually neuroscience – through my work studying the biomedical applications of organic and nanomaterials. A brief stint studying gamma oscillations in a neurophysics lab out at UCLA cemented this transition, and my interest in computational neuroscience was born.
One grad school application cycle later, I ended up at Princeton University. With a strong desire to work on mathematical and computational methods while staying “close to the experiments” (as I like to say), I quickly found my home working both in Jonathan Pillow’s Neural Coding and Computation Group and in Ilana Witten’s lab studying the Neural Circuitry of Reward Learning and Decision-Making.
PhD Research
Quantitative modeling of natural behavior and social interactions in rodents
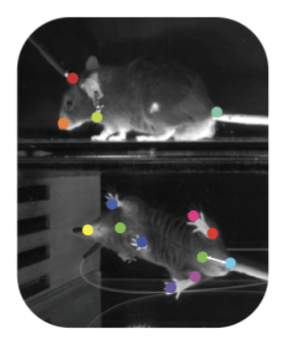
With the help of behavior tracking software like DeepLabCut and SLEAP, the quantification of natural behavior has become an increasingly popular area of study in neuroscience. However, how to best take high-dimensional measurements of behavior (such as joint positions or pixel data) and identify interpretable, low-dimensional representations (i.e. “syllables” of behavior such as walking or grooming) from that data is still an open question. This is an important line of research not only because it can help with the automated, unsupervised identification of behavior in a variety of animals, but also because it can help map these behaviors to changes in neural activity as observed in electrophysiology, calcium imaging, and other types of neural data. Latent variable models are a powerful tool for addressing these challenges, and I’m working currently working on some new methods in this area. Stay tuned!
Keywords/techniques: latent variable models, generative models, behavior quantification, unsupervised learning
Code: python (statistical modeling, data analysis, data visualization)
Spectral learning of Bernoulli linear dynamical systems (LDS) models
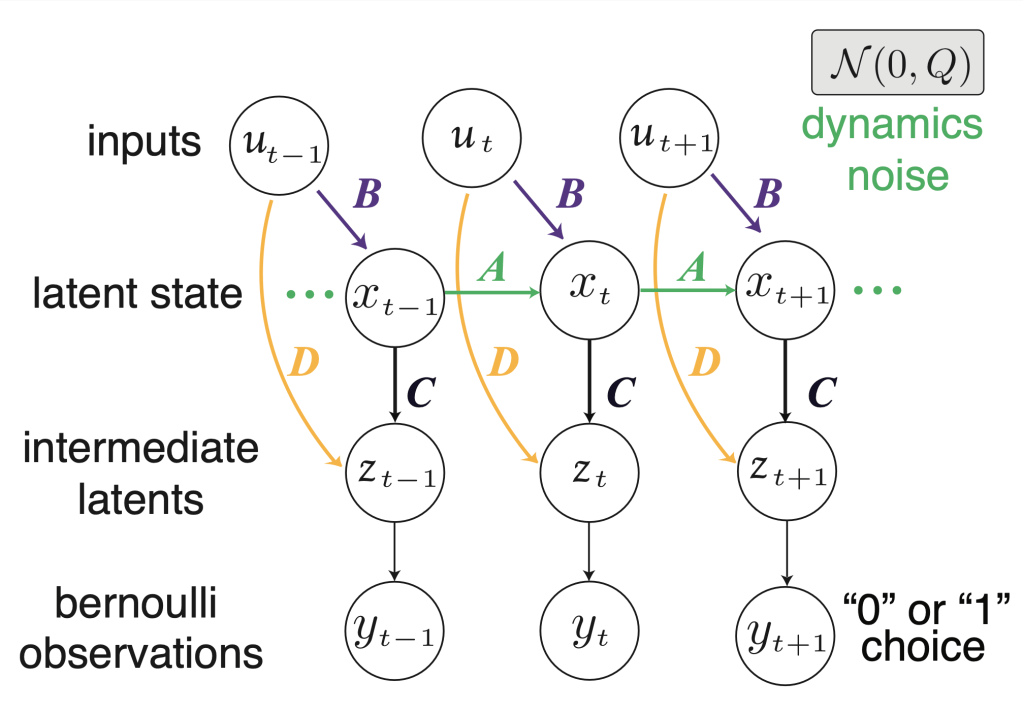
Accurately identifying the parameters of generative models like linear dynamical systems (LDS) can be a complicated process involving the use of time-consuming and computationally-intensive methods such as the expectation maximization (EM) algorithm. Therefore, fast methods for approximating these parameters — either to a satisfactory degree of accuracy or for use as intelligent initializations in optimization techniques like EM — can provide great advantages in the use of LDS models. Here, we develop a spectral learning method for fast, efficient fitting of input-driven Bernoulli LDS models that extends traditional subspace identification methods and is ideal for application to binary time-series data that are common in neuroscience, such as choice behavior in two alternative forced choice (2AFC) decision-making tasks and binned neural spike trains.
Keywords/techniques: latent variable models, generative models, linear dynamical systems, subspace identification, spectral learning, decision-making, 2AFC tasks
Code: python (public github package available here)
Latent variable modeling of the strategy-dependent role of the DMS in decision-making

When mice make decisions, do they use the same strategy trial after trial, or do they switch strategies over time? What brain regions are involved in this possible strategy-switching behavior? In this project, I applied a hybrid model consisting of a Generalized Linear Model (GLM) and a Hidden Markov Model (HMM) to behavioral data of mice performing a two alternative forced choice (2AFC) decision-making task while receiving optogenetic inhibition of either the direct (D1) or indirect (D2) pathway of the dorsomedial striatum (DMS) on a subset of trials. We found that not only do mice use different strategies when making decisions (guided by different external and behavioral variables such as the task stimulus or previous choice), but that the role of the DMS in decision-making is strategy-dependent.
Keywords/techniques: latent variable models, generative models, generalized linear models, hidden Markov models, expectation maximization algorithm, statistical modeling, decision-making, 2AFC tasks, dopamine, striatum
Code: python (public github package available here)
Undergraduate Research
Analyzing the dependence of hippocampal gamma oscillations on running speed

During undergrad, I spent a summer working at UCLA as part of an NSF-sponsored Research Experience for Undergraduates (REU). There, I analyzed local field potential (LFP) data, looking at high frequency (gamma, 30-140 Hz) oscillations in the hippocampus of rats while running mazes in both the “real world” and virtual reality (VR). The goal was to compare what type of cues (e.g. distal, self-motion, vestibular, and sensory — only the former two of which are present in VR) are responsible for modulating gamma rhythms. We also explored how changes in theta phase and phase precession could contribute to this modulation. Our results suggested that two categories of gamma oscillations – slow gamma (30-60 Hz) and fast gamma (60-140 Hz) may have distinct roles in encoding running speed, with different sensitivity to environmental cues.
Keywords/techniques: local field potential, EEG, gamma oscillations, hippocampus, signal processing, power spectral analysis, theta phase
Code: MATLAB (data analysis and visualization)
Characterizing the optoelectronic properties of nano-materials

As a physics undergraduate, I worked with Dr. Patrick Vora on a variety of projects exploring the optical and electronic properties of low-dimensional materials. This included studies focused on identifying the stoichiometry, morphology, and charge transfer properties of organic materials for use in biomedical devices; assessing the optical absorption properties of single-walled carbon nanotubes (SWNTs) when coupled with dopamine for use in real-time neurotransmitter sensors; and discovering new 2D materials (e.g. transition metal dichalcogenides, or TMDs) for use in next-generation electronics.
Keywords/techniques: Raman spectroscopy, cryogenics, microscopy, optical physics
Code: LabVIEW (instrumentation), MATLAB (data visualization), python (analysis)